적 요본 연구에서는 간혹 흰바위취(Micranthes manchuriensis)와 동일종으로 취급되기도 하는 한국 고유종 구실바위취(Saxifraga octopetala)의 분류학적 지위를 알아보고자 수행되었다. 구실바위취의 계통학적 위치와 종의 경계를 확인하기 위해 두 종의 기준표본에 대한 형태적 검토와 핵 리보솜 DNA의 ITS 지역 염기서열에 대한 집중적인 계통분석을 시행하였다. 총 65개 구실바위취 개체는 ITS 계통수에서 하나의 무리를 이루면서 Micranthes 분기군(clade)에 포함되었고, 톱바위취 및 흰바위취와 가까운 계통 유연관계를 나타내었다. 중국과 러시아에서 채집된 다수의 흰바위취 개체 역시 독자적인 분기군을 형성하면서 M. nelsoniana var. pacifica 및 M. fusca와 자매군을 이루었다. 구실바위취의 실체가 모호했던 것은 흰바위취 개체와 톱바위취의 화서를 함께 포함하고 있는 복합표본인 Wilford의 채집품(흰바위취의 기준표본)을 Nakai가 잘못 관찰하였기 때문으로 생각되었다. 구실바위취와 흰바위취의 높은 형태적 유사성에도 불구하고 이들은 지하 포복경의 특징에 있어서 차이를 보였다. ITS 계통수에서 구실바위취가 Micranthes 내에 자리하는 것으로 나타나 구실바위취는 Micranthes 내에서 종의 지위를 유지하여야 할 것으로 판단되었고, 이에 따라 신조합명(Micranthes octopetala)을 발표하였다.
AbstractThis study was conducted to address the taxonomic status of the Korean endemic species Saxifraga octopetala, which is sometimes considered conspecific to Micranthes manchuriensis. Extensive molecular phylogenetic analysis using nrITS sequences as well as morphological examination of type specimens of the two species were undertaken to ascertain the phylogenetic position and species delimitation of S. octopetala. In the resulting nrITS trees, a total of 65 accessions representing S. octopetala grouped together and nested within the Micranthes clade, exhibiting a close relationship with M. nelsoniana and M. manchuriensis. Multiple accessions of M. manchuriensis collected from China and Russia also formed a clade, showing a sister group relationship with M. nelsoniana var. pacifica and M. fusca. The ambiguous species entity of S. octopetala is thought to have originated from Nakai's misinterpretation of Wilford's collection (type specimens of M. manchuriensis), which is a complex collection including an inflorescence of M. nelsoniana. In spite of apparent morphological similarity between S. octopetala and M. manchuriensis, they differ in the presence and absence of underground stolons. The distinct position of S. octopetala within the Micranthes clade on the nrITS tree suggests that it should retain species status in Micranthes. Thus a new combination (Micranthes octopetala) is proposed.
IntroductionSince Soltis et al. (1996) reported polyphyly of Saxifraga L. sensu lato, many species within the genus have been transferred to other genera, especially to Micranthes Haw. (Gornall et al., 2000; Brouillet and Gornall, 2007; Elven and Murray, 2008; Akiyama et al., 2012; Tkach et al., 2015). Micranthes used to be classified as a subgenus or section under Saxifraga s.l., which includes approximately 400 species (Engler and Irmscher, 1916; Gornall, 1987); however, various molecular phylogenetic studies have confirmed repeatedly that the Micranthes clade is distinct from Saxifraga sensu stricto (Soltis et al., 2001, 2013; Xiang et al., 2012; Prieto et al., 2013; Deng et al. 2015; Tkach et al., 2015). Moreover, it was revealed that many genera (i.e., Chrysosplenium L., Astilbe Buch.-Ham. ex D.Don, Mukdenia Koidz., Mitella L.) have a closer phylogenetic relationship with Micranthes than with the Saxifraga s.s. clade, granting genus status of Micranthes (Soltis et al., 2001; Deng et al., 2015; Tkach et al., 2015).
The segregation of Micranthes from Saxifraga s.l. was also supported by several morphological features, such as absence of cauline leaves, seed coat with vertical ridges, and unitegmic ovule (Webb and Gornall, 1989; Brouillet and Elvander, 2009). As a result, new name combinations were made for most of the Northeast Asian taxa such as M. oblongifolia (Nakai) Gornall & H. Ohba (= S. oblongifolia Nakai), M. laciniata (Nakai & Takeda ex Nakai) S. Akiyama & H. Ohba (= S. laciniata Nakai & Takeda ex Nakai), and M. manchuriensis (Engl.) Gornall & H. Ohba (= Saxifraga manchuriensis (Engl.) Komarov) (Akiyama et al., 2012). Nevertheless, taxonomic treatment of S. octopetala Nakai, a closely related species to M. manchuriensis, has not yet been conducted.
Saxifraga octopetala is a perennial herbaceous species, endemic to Korea. Nakai (1918) described the species based on four specimens collected from the eastern part of North Korea. In South Korea, it grows near small shady creeks or on moist rock crevices in high mountain areas, with the southern limit in Sobaeksan Mt. Morphologically, S. octopetala strongly resembles M. manchuriensis, which is distributed in eastern Jilin in China and Southern Primorsky Krai in Far East Russia. In the description of S. octopetala, Nakai (1918) listed the following diagnostic characters (compared with M. manchuriensis): rhizome is not fasciculate, underground stolons present, flowers arranged more densely in panicle, number of petals is not five but eight, lengths of petals and filaments are two and three times longer than sepals, respectively, and the shape of anther is spheric (vs. oblong). However, these diagnostic characters, especially the floral ones, are known to be unuseful because of disagreement between the species description and field observation. Thus sometimes S. octopetala is considered conspecific to M. manchuriensis (Chang et al., 2014).
This study was conducted to ascertain the phylogenetic position and species delimitation of S. octopetala within Micranthes. To achieve these goals, we performed phylogenetic analysis of nrITS (Internal Transcribed Spacer regions of nuclear ribosomal DNA) sequences, which have proven to be very useful for delimiting species boundaries and elucidating phylogenetic relationships of various genera in Saxifragaceae (Okuyama and Kato, 2009; Kim and Kim, 2011; Xiang et al., 2012; Gao et al., 2015; Tkach et al., 2015). In addition, the type specimens of S. octopetala and M. manchuriensis were reexamined to check the diagnostic morphological characters of the taxa. Based on the results of the phylogenetic and morphological examinations, a new name combination (Micranthes octopetala Y.I. Kim & Y.D. Kim) for S. octopetala was proposed.
Materials and MethodsTaxon samplingA total of 114 nrITS sequences representing 20 taxa of Micranthes and three outgroup species were included for phylogenetic analysis (Table 1). The ingroup taxa were chosen to represent eight major sections of Micranthes (Tkach et al., 2015). Three outgroup taxa (Chrysosplenium alternifolium L., Chrysosplenium valdepilosum (Ohwi) S.H. Kang & J.W. Han, and Peltoboykinia tellimoides (Maxim.) H. Hara) were selected based on previous phylogenetic studies (Xiang et al., 2012; Deng et al., 2015). Among the 114 nrITS sequences, 93 sequences were newly generated in this study: 65 accessions of M. octopetala from four populations in South Korea; 24 accessions of M. manchuriensis from Jilin, China and Vladivostok, Russia; and four accessions of M. nelsoniana var. aestivalis (Fischer & C.A. Meyer) Gornall & H. Ohba from Jilin, China (Table 3). All the leaves used for the DNA sequencing were collected in the flowering period to minimize the possibility of misidentification. Voucher specimens representing each population of the above three species were deposited at KB: M. octopetala −− Daeseongsan Mt. (KYI2013106, KYI2013107), Hwaaksan Mt. (KYI2013036, KYI2013038), Chiaksan Mt. (KYI2013128, KYI2013129), Sobaeksan Mt. (KYI2013141, KYI2013160); M. manchuriensis −− Jilin (KYI2013168, KYI2013169), Vladivostok (KYI2013223, KYI2013224); and M. nelsoniana var. aestivalis −− Jilin (KYI2013185, KYI2013202). The nrITS sequences of other Micranthes taxa and three outgroup species were downloaded from GenBank (Table 1).
DNA extraction, PCR, and phylogenetic analysisDNA was extracted from dried leaves using DNeasy Plant Mini Kits (Qiagen, Germany) in accordance with the manufacturer's instructions. PCR amplification of the nrITS region was carried out in a total reaction volume of 50 μL containing 5 μL of Takara 10X Ex-Taq buffer, 4 μL of 2.5 mM dNTPs, 0.5 μL each of forward and reverse primers, 0.25 μL of 5 U/μL Ex-Taq polymerase, 10−20 ng of template DNA, and distilled water up to the final volume. The thermocycling profile consisted of an initial denaturation step at 95oC for 5 min; followed by 30 cycles of 1 min at 95oC, 1 min at 57oC, and 1 min at 72oC; and a final extension step of 10 min at 72oC. The products were purified with a QIAqiuck PCR purification kit (Qiagen, Germany) according to the manufacturer's instructions. Purified double-stranded PCR products were used for determining the DNA sequence of the nrITS region using the automatic DNA sequencer ABI PRISM 377 (PE Applied Biosystems). The nrITS regions were amplified and sequenced using primers designed by White et al. (1990), except for ITS1, which differed by the two italicized bases (5-GGA AGG AGA AGT CGT AAC AAG G-3).
DNA sequences were aligned with MAFFT (Katoh et al., 2002). The alignment was further examined and slightly edited manually as needed. Gaps introduced from the alignment were treated as missing characters in subsequent analyses. Phylogenetic analyses were performed using Maximum Parsimony (MP), Maximum Likelihood (ML), and Bayesian inference (BI) methods. MP and ML analyses were conducted in PAUP* ver.4.0b10 (Swofford, 2002). BI was conducted with MrBayes ver. 3.1.2 (Ronquist and Huelsenbeck, 2003). For the MP and ML phlyogenetic analyses, heuristic searches with the MULTREES option, ten random entries of taxa, and TBR (Tree Bisection-Reconnection) branch swapping were selected. TVM + G model was applied for ML and BI analyses as the best fit evolutionary model, which was calculated by using jModelTest 2.1.7 (Darriba et al., 2012). Bootstrap analyses (Felsenstein, 1985) with 1000 replicates were conducted to evaluate the degree of support for given clades for ML and MP analysis using the same options as tree search. In BI analysis, each Markov chain was started from a random tree and run for 2,000,000 generations, sampling trees every 200 generations, with four independent chains running simultaneously. After discarding burn-in samples (initial 2500 trees), the remaining samples were retained for the construction of a final tree with posterior probabilities for given clades. The strict consensus tree of 24 equally parsimonious trees (Fig. 2) and a BI tree (Fig. 3) were presented for the discussion of phylogenetic relationships.
MorphologySpecimens of Micranthes nelsoniana var. aestivalis, M. manchuriensis, and M. octopetala collected by the authors as well as type specimens of M. octopetala (= S. octotpetala) deposited in TI and M. manchuriensis were used to verify the diagnostic characters of M. octopetala. Two type specimens of M. manchuriensis (= Saxifraga manchuriensis), which are assumed to have been referred by Nakai (1918), were observed using the high resolution digital images provided by K (http://apps.kew.org/herbcat/getImage.do?imageBarcode=K000618434) and HUH (http://kiki.huh.harvard.edu/databases/specimen_search.php?mode=details&id=255501). The image of the specimen (isotype) provided by HUH for the discussion of taxonomic identity of S. octopetala is presented in Fig. 4.
ResultsSequence variation and ribotypesThe ribotypes (nrITS sequence types) found in each population of M. octopetala, M. manchuriensis, and M. nelsoniana var. aestivalis are listed in Table 2. All 65 accessions in M. octopetala have nrITS sequence size of 664 bp. They were clustered into 6 ribotypes according to variation observed in the 3 variable nucleotide sites. Among the four populations, Chiaksan Mt. exhibited the highest diversity in nrITS sequence types (Table 3). The sequences of 24 accessions of M. manchuriensis were 657 to 658 bp in length and were grouped into 5 ribotypes based on variations in 4 nucleotide sites (Table 2). In M. manchuriensis, higher ribotype diversity was observed in the Vladivostok population (Table 3). The sequence lengths of M. nelsoniana var. aestivalis were all 659 bp. They were classified into 2 ribotypes by the variation in 3 nucleotide sites: type 1 and 2 with three and one accession, respectively (Table 3).
The sequence lengths of the nrITS regions of the examined taxa ranged from 637 bp (Micranthes brachypetala (Malyschev) Gornall & H. Ohba) to 669 bp (Peltoboykinia tellimoides (Maxim.) H. Hara). The aligned nrITS region was 789 bp long including 358 variable sites, among which 272 sites were parsimony informative. M. octopetala, M. manchuriensis, and M. nelsoniana var. aestivalis showed a total of 49 sites of sequence variation, which were useful in distinguishing each taxon. The amount of nrITS sequence difference between M. octopetala and M. manchuriensis ranged from 18 to 26 bp, depending on accessions (or their ribotypes) compared.
Phylogenetic analysesTwenty-four equally parsimonious trees were obtained as a result of MP analysis. The length of the phylogenetic tree was 806, and CI (Consistency Index) and RI (Retention Index) were 0.6725 and 0.7604, respectively. The strict consensus tree is shown in Fig. 2. The BI tree (Fig. 3) exhibited almost identical topology to the strict consensus tree. In both trees, the Micranthes clade was evident with high bootstrap values (98/ 100 in MP/ML analysis) and posterior probability (1.00). S. octopetala grouped with M. manchuriensis, M. nelsoniana, and M. fusca within the section Rotundifoliatae (A.M. Johnson) Tkach of the Micranthes clade, which was supported by bootstrap values (97/79 in MP/ML analysis) and posterior probability (1.00).
Sixty-five accessions of S. octopetala formed a robust clade with high bootstrap values (100/96 in MP/ML analysis) and posterior probability (1.00). All 24 accessions of M. manchuriensis collected from China and Russia also formed a strong monophyletic group with high bootstrap values (100/96 in MP/ML analysis) and posterior probability (1.00). In both trees, M. manchuriensis showed a sister group relationship with a clade consisting of M. nelsoniana var. pacifica and M. fusca (Maxim.) S. Akiyama & H. Ohba (bootstrap values of 78/81 in MP/ML analysis; posterior probability of 1.00), making the M. nelsoniana complex non-monophyletic.
Morphological examinationAs a result of examining four type specimens of Saxifraga octopetala, scanned images of Micranthes manchuriensis types, and specimens collected by the authors from 6 populations in Korea, China, and Russia, no definite morphological differences in the leaf shape or floral features were found between the two taxa. Careful observation on type materials and other specimens revealed that both S. octopetala and M. manchuriensis have flowers with 8 (or sometimes 9) petals. It was confirmed that the type specimen of M. manchuriensis deposited in HUH was a complex collection consisting of a M. manchuriensis individual and an inflorescence assumed to be from another species of Micranthes, probably M. nelsoniana var. aestivalis, which has a loose inflorescence and flowers with 5 petals (Fig. 4). The presence of underground stolons, a diagnostic character state of S. octopetala suggested by Nakai (1918), was not clearly seen in the type specimens although they were observed from the specimens collected in this study.
DiscussionDelimiting or recognizing species by using DNA data and subsequent reexamination of non-molecular traits is an active area of systematic research (Lpez et al., 2007; Wiens, 2007; Okuyama and Kato, 2009; Ruiz-Sanchez and Sosa, 2010; Efimov, 2013; Boatwright et al., 2015). The nrITS sequence data utilized for this study proved to be very useful in delimiting the Korean endemic Saxifraga octopetala (= Micranthes octopetala) from M. manchuriensis and other related taxa such as M. nelsoniana var. aestivalis (Table 2). Considering the strong morphological similarities between S. octopetala and M. manchuriensis, an nrITS sequence difference of 18 to 26 bp between the two taxa is a noteworthy result. This suggests that substantial genetic divergence has occurred between these lineages. The resulting nrITS trees (Figs. 2, 3) also indicate that S. octopetala is a well segregated lineage that merits a distinct species status within Micranthes. Moreover, merging S. octopetala with M. manchuriensis is phylogenetically inappropriate as the latter species shows a closer relationship with M. nelsoniana var. pacifica and M. fusca than with S. octopetala (Figs. 2, 3).
Although all other species of Saxifraga s.l. distributed in Korea have been transferred to Micranthes (Akiyama et al., 2012), the taxonomic treatment of S. octopetala has been neglected, probably because of its limited distribution and ambiguous species entity, especially in relation to the unclear species delimitation between the species and M. manchuriensis. The morphological difference between the two species was not apparent because most of the diagnostic characters of S. octopetala (especially reproductive ones) proposed by Nakai (1918) were mistaken, making them inapplicable for species identification. For example, both S. octopetala and M. manchuriensis were confirmed to have 8 (sometimes 9) petals, yet Nakai (1918) stated that the latter species has 5 petals.
The reason why Nakai (1918) recognized the petal number of M. manchuriensis as 5 and not 8 needs to be considered. By paying attention to the mis-mounted inflorescence of other species (probably, M. nelsoniana var. aestivalis; Fig. 4), Nakai (1918) seems to have misinterpreted the floral characteristics of M. manchuriensis. Neither Engler (1872) nor Komarov (1903) mentioned the number of petals in their description or taxonomic treatment of M. manchuriensis. It is highly probable that Nakai (1918) determined the floral features (number of petals, stamen characteristics, etc.) of M. manchuriensis by examining the mis-mounted loose inflorescence, which was relatively easier for observation of flowers, rather than the inflorescence of the genuine individual with congested flowers. Therefore, it can be inferred that the diagnostic characters of the floral part listed in Nakai (1918) are probably those of M. nelsoniana var. aestivalis.
It is intriguing that M. manchuriensis showed a closer phylogenetic relationship with M. nelsoniana var. pacifica and M. fusca than with S. octopetala, which sometimes is considered conspecific to M. manchuriensis (Chang et al., 2014). It should be emphasized that this result is based on examination of multiple (up to 65) accessions representing each taxon and on careful collection of leaves (from flowering individuals) for DNA analysis. Identical topology and relatively high statistical supports for the relationship (Figs. 2, 3) suggest that the discordance between phylogenetic affinity and morphological similarity is largely irrelevant to the employed phylogenetic analysis methods.
Symplesiomorphy of M. manchuriensis and S. octopetala might be the cause of the morphological similarity. In other words, morphological traits of their ancestor seem to have continued to exist in both species even after genetic differentiation, leading to the apparent morphological similarity. The evolutionary phenomenon known as morphological stasis (Larson, 1989) has been reported in several plant groups including Aralia sect. Dimorphanthus (Wen, 2000), Liquidambar (Hoey and Parks 1991; Shi et al., 1998), Magnolia sect. Rytidospermum (Qiu et al., 1995), Osmorhiza (Wen et al., 2002), and Phryma (Nie et al., 2006). Unlike the above cases, which dealt with morphological stasis of taxa showing intercontinental disjunctive distribution, our result is noteworthy as it is observed from taxa distributed adjacent to each other in northeast Asia.
The sister group relationship between M. manchuriensis and the clade consisting of M. nelsoniana var. pacifica and M. fusca poses another taxonomic issue: non-monophyly of M. nelsoniana. It is known that many intraspecific taxa of M. nelsoniana are widely distributed in cool-temperate regions in the northern hemisphere (Brouillet and Elvander, 2009). Non-monophyly of the species complex was also observed in a previous study (Tkach et al., 2015). As with other species complexes, many of the intraspecific taxa have often been treated as distinct species. For example, M. nelsoniana var. pacifica has been treated as Saxifraga pacifica (Hultdèn) Zhmylev & V.V. Petrovsky. Thus, the current taxonomy of the M. nelsoniana complex needs to be tested by more detailed phylogenetic and morphological reexamination using extensive sampling encompassing all known taxa in the species complex in the northern hemisphere. Sufficient taxon sampling will be important because there are reports that the relationship of ingroups often shows different patterns if there is an insufficient number of taxonomic groups representing various lineages in the phylogenetic analysis (Zwickl and Hillis, 2002; Jockusch et al., 2015).
Meanwhile, the presence or absence of underground stolons would be a good character to distinguish S. octopetala and M. manchuriensis. Also, the rhizome of M. manchuriensis tends to be thicker than that of S. octopetala, although this feature is less consistent than the feature of underground stolons. Our study suggests that vegetative characters, including underground parts, are more useful than the known floral ones in morphological delimitation of these taxa. The characteristics of vegetative parts, such as growth pattern of sterile branches, have been verified as important in recognizing new cryptic species in the neighboring genus Chrysosplenium (Han et al., 2011; Kim and Kim, 2014; Kim 2015). The morphological difference between the two taxa should be explored further through a detailed comparative study of additional characters such as fruit and seed morphology, which have not been examined thus far. Based on the result of molecular phylogenetic analysis and morphological difference (i.e., presence or absence of underground stolons) between S. octopetala and M. manchuriensis, a new combination for S. octopetala is presented below.
Taxonomic Treatment
Micranthes octopetala (Nakai) Y. I. Kim & Y. D. Kim, comb. nov.
Basionym: Saxifraga octopetala Nakai, Bot. Mag. (Tokyo) 32: 230. 1918. TYPE: Korea. Chagang-do, Taeheungri− Sanyang, 21 July 1914 T. Nakai 6439 (lectotype, TI!); Hamgyong-bukto, Ranan, 1913, I. Ono s.n. (syntype, TI!); Kangwon-do, Ouensan, July 1906, U.J. Faurie 377 (syntype, TI!); Rason, Solsinsan, 28 August 1914, T. Nakai 6450 (syntype, TI!).
Herbs, perennial, to 15−30 cm, with underground stolons; rhizomes short, transversely elongating. Flowering stems erect, leafless, glandular, villous. Leaves 1−4, basal; stipules absent; petiole 4.7−13 cm long, sparsely glandular piliferous distally; blade reniform to orbicular-cordate, 3.8−8.5 cm long, 4.2− 10.3 cm wide, base reniform, apex rounded, margin 23−32- crenate, abaxially and marginally strigulose, palmately veined. Inflorescence paniculate, compact, many flowered, 3−9.3 cm long; bracts lanceolate or linear, 4−7 mm long; pedicels glandular hairy. Flower: bisexual, radially symmetric; sepals 8 (rarely 9), reflexed, sublanceolate, 2.0−2.5 mm long, 0.4− 0.7 mm wide, apex subobtuse, glandular hairy on abaxial surface, 1-veined, margin glandular ciliate; petals 8 (rarely 9), white, oblong-oblanceolate, ca. 4 mm long, ca. 1 mm wide, apex subobtuse or rounded or minutely retuse, 1-veined; stamens 15−17, 3.6−5.8 mm long; filaments white, slightly clavate; anthers spheric to oblong, orange; carpels 2 (or 3), superior, united only at base, white, conical; styles 2 (or 3), erect or slightly curved, 3.3−3.4 mm long; stigma capitate. Capsules erect, divergent, 4.5−6.0 mm long.
Korean Name: Gu-sil-ba-wi-chui (구실바위취)
Flowering: Jun. to Jul.
Distribution: Korea (endemic).
Korea: N Korea (HB, HN), GG, GW, CB. Wet places along shady creeks or on rock crevices in high mountain areas.
ACKNOWLEDGMENTSWe thank Dr. Kanchi Gandhi for the helpful comments on the taxonomy part of the manuscript. We also give our thanks to Prof. Gorovoy for helping during our field trip in the Primorsky area. This study was supported by a grant titled “The Genetic Evaluation of Important Biological Resources 2013” from the National Institute of Biological Resources under the Ministry of Environment, Republic of Korea.
Fig 1.A map showing the populations of Micranthes octopetala (= Saxifraga octopetala), M. manchuriensis, and M. nelsoniana var. aestivalis examined in this study. 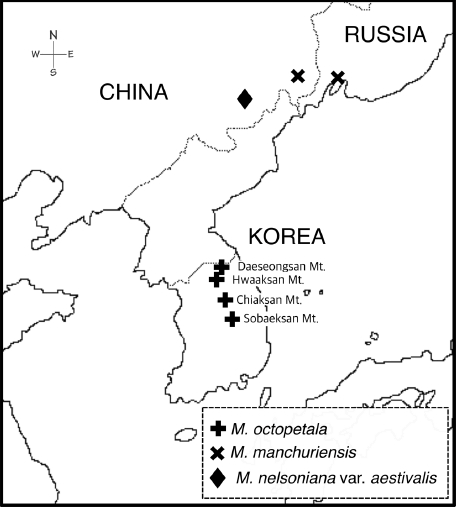
Fig 2.Strict consensus trees of 24 equally parsimonious trees of Micranthes based on nrITS sequences. Asterisk mark indicates a taxon that was newly sequenced in this study. Numbers after a taxon name denote nrITS sequence types (ribotypes; refer to Tables 2 and 3 for each ribotype and its distribution among the populations). Numbers above branches are bootstrap values of MP/ML analyses. 
Fig 3.A Bayesian inference tree of Micranthes based on nrITS sequences. Asterisk mark indicates a taxon that was newly sequenced in this study. Numbers after a taxon name denote nrITS sequence types (ribotypes; refer to Tables 2 and 3 for each ribotype and its distribution among the populations). Numbers above or below branches are posterior probabilities. 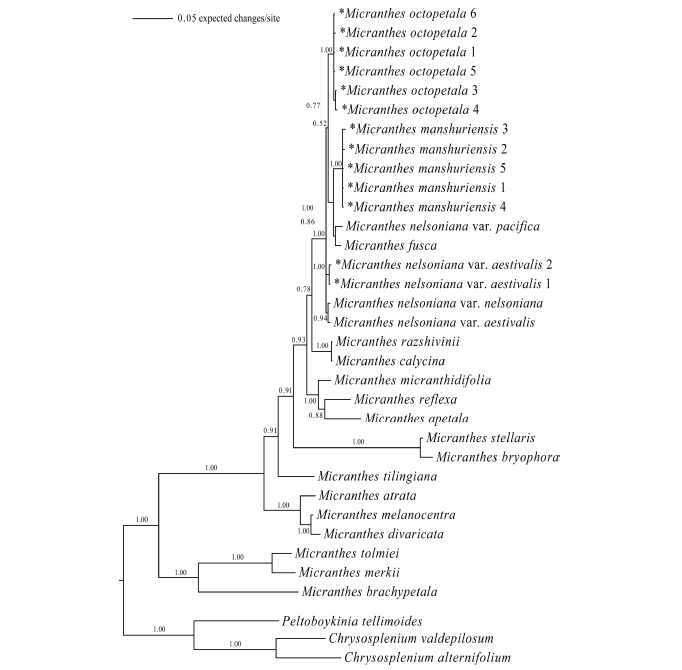
Fig 4.Type specimen of Micranthes manchuriensis deposited in HUH. This is a complex collection object (2 items with 1 preparation). Inside the broken line is an inflorescence, assumed to be from another species of Micranthes, probably of M. nelsoniana var. aestivalis. 
Table 1.Sequence information on Micranthes and three outgroup taxa (Chrysosplenium alternifolium, C. valdepilosum, and Peltoboykinia tellimoides) examined in this study.
Table 2.Sequence variation among the nrITS sequence types (ribotypes) obtained from 65 accessions representing Micranthes octopetala, M. manchuriensis, and M. nelsoniana var. aestivalis. Gray shades indicate distinct bases or gaps segregating each taxon from other two taxa. “-” denotes a gap. R, W, and Y are letter abbreviations for mixed base positions for A and G, A and T, and C and T, respectively. See Table 3 for the distribution and frequency of each ribotype in each population. Table 3.The nrITS sequence types (ribotypes) observed from Micranthes octopetala, M. manchuriensis, and M. nelsoniana var. aestivalis populations, their GenBank number, and populations with the number of accessions exhibiting each ribotype. Literature CitedAkiyama, S. Gornall, R. J and Ohba, H. 2012. Asiatic species of the genus Micranthes Haw. (Saxifragaceae). Journal of Japanese Botany 87: 236-240.
Boatwright, J. S. Maurin, O and van der Bank, M. 2015. Phylogenetic position of Madagascan species of Acacia s.l. and new combinations in Senegalia and Vachellia (Fabaceae, Mimosoideae, Acacieae). Botanical Journal of the Linnean Society 10.1111/boj.12320.
Brouillet, L and R., Gornall. 2007. New combinations in Micranthes (A segregate of Saxifraga, Saxifragaceae) in North America. Journal of the Botanical Research Institute of Texas 1: 1019-1022.
Brouillet, L and Elvander, P. E. 2009. Micranthes Haw. In Flora of North America, Vol. 8. Editorial Committee (ed.), Oxford University Press, New York. 49-70.
Chang, C.-S. Kim, H and Chang, K. S. 2014. Provisional checklist of vascular plants for the Korean Peninsula Flora (KPF) (version 1.0). 660.
Darriba, D. Taboada, G. L. Doallo, R and Posada, D. 2012. jModelTest 2: more models, new heuristics and parallel computing. Nature Methods 9: 772.
Deng, J.-b. Drew, B. T. Mavrodive, E. V. Gitzendanner, M. A. Soltis, P. S and Soltis, D. E. 2015. Phylogeny, divergence times, and historical biogeography of the angiosperm family Saxifragaceae. Molecular Phylogenetics and Evolution 83: 86-98.
Efimov, P. G. 2013. Sibling species of fragrant orchids (Gymnadenia: Orchidaceae, Magnoliophyta) in Russia. Russian Journal of Genetics 49: 299-309.
Elven, R and Murray, D. F. 2008. New combinations in the Panarctic vascular flora. Journal of the Botanical Research Institute of Texas 2: 433-446.
Engler, A. 1872. Monographie der Gattung Saxifraga L., mit besonderer Berücksichtigung der geographischen Verhältnisse, von A. Engler. Mit einer lithographirten Karte. 139 pp.
Engler, A and Irmscher, E. 1916. Section Boraphila Engler. In Das Pflanzenreich, Vol. 67. Saxifragaceae - Saxifraga. Engler, A (ed.), Engelmann, Leipzig. 5-89.
Felsenstein, J. 1985. Confidence limits on phylogenies: an approach using bootstrap. Evolution 39: 783-791.
Gao, Q-B. Li, Y.-H. Gornall, R. J. Zhang, Z.-X. Zhang, F.-Q. Xing, R. Fu, P.-C. Wang, J.-L. Liu, H.-R. Tian, Z.-Z and Chen, S.-L. 2015. Phylogeny and speciation in Saxifraga sect. Ciliatae (Saxifragaceae): Evidence from psbA-trnH, trnL-F and ITS sequences.. Taxon 64: 703-713.
Gornall, RJ. 1987. An outline of a revised classification of Saxifraga L. Botanical Journal of the Linnaean Society 95: 273-292.
Gornall, RJ. Ohba, H and Pan, J. 2000. New taxa, names, and combinations in Saxifraga (Saxifragaceae) for the Flora of China. Novon 10: 375-377.
Han, JW. Yang, S. G. Kim, H. J. Jang, C. G. Park, J. M and Kang, S. H. 2011. Phylogenetic study of Korean Chrysosplenium based on nrDNA ITS sequences. Korean Journal of Plant Resources 24: 358-369.
Hoey, M. T and Parks, C. R. 1991. Isozyme divergence between eastern Asian, North American and Turkish species of Liquidambar (Hamamelidaceae). American Journal of Botany 78: 938-947.
Jockusch, EL. Martínez-Solano, I and Timpe, E. K. 2015. The effects of inference method, population sampling, and gene sampling on species tree inferences: an empirical study in slender salamanders (Plethodontidae: Batrachoseps). Systematic Biology 64: 66-83.
Katoh, K. Misawa, K. Kuma, K.-I and Miyata, T. 2002. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Research 30: 3059-3066.
Kim, Y.-I and Kim, Y.-D. 2011. Molecular systematic study of Chrysosplenium series Pilosa (Saxifragaceae) in Korea. Journal of Plant Biology 54: 396-401.
Kim, Y.-I and Y.-D., Kim. 2014. Chrysosplenium aureobracteatum (Saxifragaceae), a new species from South Korea. Novon 23: 432-436.
Kim, Y.-I. 2015. Systematic studies of Chrysosplenium L. series Pilosa Maxim. (Saxifragaceae). Ph.D. Dissertation, Hallym University, S; Korea. (in Korean).
Komarov, V. L. 1903. Flora Manshuriae t. II. pars 1. Trudy Imperatorskago S.-Peterburgskago Botanicheskago sada 22: 1-452.
Larson, A. 1989. The relationship between speciation and morphological evolution. In Speciation and its Consequences. Otte, D. Endler, J. A (eds.), Sinauer Assoc. Sunderland. Massachusetts, 579-598.
López, H. Contreras-Díaz, H. G. Oromí, P and Juan, C. 2007. Delimiting species boundaries for endangered Canary Island grasshoppers based on DNA sequence data. Conservation Genetics 8: 587-598.
Nie, Z-L. Sun, H. Beardsley, P. M. Olmstead, R. G and Wen, J. 2006. Evolution of biogeographic disjunction between eastern Asia and eastern North America in Phryma (Phrymaceae). American Journal of Botany 93: 1343-1356.
Okuyama, Y and Kato, M. 2009. Unveiling cryptic species diversity of flowering plants: successful biological species identification of Asian Mitella using nuclear ribosomal DNA sequences. BMC Evolutionary Biology 9: 105.
Prieto, J. A. F. Arjona, J. M. Sanna, M. Pérez, R and Cires, E. 2013. Phylogeny and systematics of Micranthes (Saxifragaceae): an appraisal in European territories. Journal of Plant Research 126: 605-611.
Qiu, YL. Parks, C. R and Chase, W. 1995. Molecular divergence in the eastern Asia–eastern North America disjunct section Rytidospermum of Magnolia (Magnoliaceae). American Journal of Botany 82: 1589-1598.
Ronquist, F and Huelsenbeck, J. P. 2003. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19: 1572-1574.
Ruiz-Sanchez, E and Sosa, V. 2010. Delimiting species boundaries within the Neotropical bamboo Otatea (Poaceae: Bambusoideae) using molecular, morphological and ecological data. Molecular Phylogenetics and Evolution 54: 344-356.
Shi, S. H. Chang, H. T. Chen, Q. Y. Qu, L. H and Wen, J. 1998. Phylogeny of the Hamamelidaceae based on the ITS sequences of nuclear ribosomal DNA. Biochemical Systematics and Ecology 26: 55-69.
Soltis, DE. Johnson, L. A and Looney, C. 1996. Discordance between ITS and chloroplast topologies in the Boykinia group (Saxifragaceae). Systematic Botany 21: 169-185.
Soltis, DE. Kuzoff, R. K. Mort, M. E. Zanis, M. Fishbein, M. Hufford, L. Koontz, J and Arroyo, M. K. 2001. Elucidating deeplevel phylogenetic relationships in Saxifragaceae using sequences for six chloroplastic and nuclear DNA regions. Annals of the Missouri Botanical Garden 88: 669-693.
Soltis, D. E. Mort, M. M. Latvis, M. Mavrodiev, E. V. O’Meara, B. C. Soltis, P. S. Burleigh, J. G and Rubio De Casas, R. 2013. Phylogenetic relationships and character evolution analysis of Saxifragales using a supermatrix approach. American Journal of Botany 100: 916-929.
Swofford, DL. 2002. PAUP* Phylogenetic Analysis Using Parsimony (* and other method), ver. 4.0b10. Sinauer Associates, Sunderland, MA.
Tkach, N. Röser, M and Hoffmann, M. H. 2015. Molecular phylogenetics, character evolution and systematics of the genus Micranthes (Saxifragaceae). Botanical Journal of the Linnean Society 178: 47-66.
Webb, D. A and Gornall, R. J. 1989. Saxifrages of Europe: with notes on African, American and some Asiatic species. Christopher Helm, London.
Wen, J. 2000. Internal transcribed spacer phylogeny of the Asian and eastern North American disjunct Aralia sect. Dimorphanthus (Araliaceae) and its biogeographic implications. International Journal of Plant Sciences 161: 959-200.
Wen, J. Lowry, II. Walck, J. L and Yoo, K. O. 2002. Phylogenetic and biogeographic diversifications in Osmorhiza (Apiaceae). Annals of the Missouri Botanical Garden 89: 414-428.
White, TJ. Bruns, T. Lee, S and Taylar, J. 1990. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR protocols: A guide to methods and application. Innis, M. Glfand, D. Sninsky, J. White, T (eds.), Academic Press; San Diego. 315-322.
Xiang, C.-L. Gitzendanner, M. A. Soltis, D. E. Peng, H and Lei, L.-G. 2012. Phylogenetic placement of the enigmatic and critically endangered genus Saniculiphyllum (Saxifragaceae) inferred from combined analysis of plastid and nuclear DNA sequences. Molecular Phylogenetics and Evolution 64: 357-367.
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||