KANG and KIM: New diploid populations of Chrysanthemum indicum L. (Asteraceae) from Korea
Abstract
Chrysanthemum indicum (Asteraceae) is a perennial plant belonging to the genus Chrysanthemum. The basic chromosome number of Chrysanthemum sensu stricto is x = 9, and it consists of a series of polyploids ranging from diploid to decaploid. However, C. indicum, which occurs in Korea, is known to consist of only tetraploids, except for two diploid populations that are sympatric with C. zawadskii and C. boreale. During the collection of plant materials as part of a study to ascertain the diversity of Chrysanthemum in Korea, we found new diploid populations (2n = 18) of C. indicum in the southern region of Korea and describe them here in detail.
Keywords: Chrysanthemum indicum, diploid, tetraploid, chromosome number
The genus Chrysanthemum (subfamily Asteroideae of Asteraceae) comprises 41 recognized species, mainly distributed across East Asia including Korea, China, and Japan ( Bailey and Bailey, 1976; Ohwi, 1984; Fu et al., 2005; Oberprieler et al. 2007). It is characterized by the obovoid and generally mucilaginous cypselae without pappus and involucral bracts with dark brown margins ( Bremer and Humphries, 1993; Lee, 2006; Zhao et al., 2009). Historically the genus Chrysanthemum distributed in Asia was especially separated from Chrysanthemum s. l. of the broad sense by Kiramura (1940), and was recognized as an independent genus Dendranthema (DC.) Des Moul. of the narrow sense ( Bremer and Humphries, 1993). However, it was changed again and back to the Chrysanthemum ( Trehane, 1995; Nicolson, 1999). Chrysanthemum indicum L. is a perennial plant and it has commonly yellow ray flower as well as C. boreale.
Kim and Tobe (2009) had reported that the leaf shape of C. boreale and C. indicum were similar to each other though the thickness were slightly different. However, Jeong (2011) and Song et al. (2012) found a wide variation of leaf shape within both species and sometimes it was difficult to distinguish each species because of overlapped variation. They suggested that an adaptation to their native habitats leads the similar external feature appearance between both species. Although it is obviously difficult to distinguish these closely related species to each other clearly, the head flower size has been regarded as one of key characters to classify two species. According to Lee (2003), the C. boreale’s head flower is 1.5 cm and the C. indicum is 2.5 cm in diameter. Kim et al. (2003) described that C. boreale’ head flowers was less than 1.5 cm and C. indicum was 1.5–2.5 cm in diameter. Therefore, for a long time, they have been accepted the diagnostic characters for classifying both species.
Polyploidization has played an important role in plant evolution and speciation, and it is prevalent in the genus Chrysanthemum ( Dowrick, 1952; Shimotomai et al., 1956; Tanaka, 1959, 1960; Nakata and Tanaka, 1987; Tsukaya, 2002; Kim et al., 2003). From the accumulated data of cytological studies for long time, it was clear that the basic chromosome number of the genus Chrysanthemum is x = 9 and it has a successive ploids in the species level ( Grant, 1981; Oberprieler et al. 2007). For the example, C. zawadskii has 2 x, 4 x, 6 x, 8 x, and 10 x ( Nakata and Kumagai, 1999; Kim et al., 2004), C. indicum has 2 x, 4 x, and 6 x ( Lee and Oh, 1976; Nakata et al. 1987; Taniguchi 1987; Du et al., 1989). Since Lee and Oh (1976) found two cytotypes of yellow-flowered wild chrysanthemum in Korea, one is C. boreale which has numerous small heads and 18 chromosomes and the other is C. indicum which has a few larger heads of 1.5–2.3 cm in diameter and 36 chromosomes in the mitotic cells. The information of the C. indicum (2 n = 36) population reported by Lee and Oh (1976) is not detailed and is known as a total of 59 populations. C. indicum had been known to be composed of only tetraploid population in Korea until the sympatric diploid populations ( Fig. 1, Table 1) were reported by Kim et al. (2003). For the purpose of understanding the diversity in the genus Chrysanthemum, we re-investigated the ploidy of C. indicum populations and found new diploid populations are widely distributed in Korea. Therefore, we described the details of the population information in the present study for the advanced research.
Materials and Methods
Plant materials
A total of 167 living individuals were collected from seventeen populations of C. indicum ( Fig. 1, Table 1) which are native and composed of over 20 individuals, and identified by external morphological characters of the head size and color. After collecting from the national habitats, they were managed for sufficient water supply in the greenhouse of Chungbuk National University. The voucher specimens were deposited in the herbarium of Department of Forest Science, Chungbuk National University (CBNU).
Root fixation
According to Kim et al. (2003), fresh root tips were collected from the planted living materials and pre-treated with 0.002 M 8-hydroxyquinolin solution for 2 h at 20°C. It was then washed with distilled water, and fixed with Carnoy’s solution (45% acetic acid:99% ethanol = 1:3) for 30 min on ice. And they were stored in 70% cold ethanol at −20°C for the mitotic chromosome observation.
Mitotic chromosome observations
To observe the mitotic metaphase chromosomes, the squash method was performed with the same condition of Kim et al.’s study (2003). The root tips were dissociated at 65°C for 1 min pre-heated 1 N HCl solution and stained by 1% aceto-orcein solution.
It was repeated at least five times per individual, and the chromosome numbers were observed under the optical microscope (Olympus BX50, Tokyo, Japan) to verify their chromosome number.
Results and Discussion
We found that C. indicum showed various leaf shapes ( Fig. 2) even though they were identified as C. indicum based on the head flower size and leaf shape. Based on the observation of mitotic metaphase chromosome, we found that new diploid populations of C. indicum are distributed in Korea. Out of 17 populations which were investigated in the present study, 13 populations were diploid of 2 n = 18 (pop No. 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, and 13) and 4 populations were tetraploid of 2 n = 36 (pop No. 14, 15, 16, and 17) ( Fig. 3, Table 1). These newly reported diploid populations of C. indicum could be used as an important material for understanding the evolution patterns of C. indicum in Korea. The most important mechanisms for understanding plant evolution are hybridization and polyploidization ( Dowrick, 1952; Shimotomai et al., 1956; Tanaka, 1959, 1960; Lee, 1975; Nakata and Tanaka, 1987; Nakata et al., 1987; Hotta et al., 1996; Tsukaya, 2002; Kim et al., 2003). C. zawadskii complex has been reported from 2 x (2 n = 18) to 10 x (2 n = 90), whereas C. indicum is composed of diploid and tetraploid, and C. boreale is specifically known only diploid ( Lee and Oh, 1976; Nakata et al. 1987; Taniguchi 1987; Du et al. 1989; Nakata and Kumagai 1999; Kim et al. 2004; Kim et al. 2008). From the result, C. indicum should be recognized as a species complex as well as C. zawadskii complex composed of different ploid level of diploid and tetraploid for the further study. It needs extensive cytological studies of C. indicum for clarifying the diversification mechanism and relationship among the populations. Also it is suggested the probability that the expression of polyploidization is differ depend on the species. In addition, the presence of intermediate type of both species, C. indicum and C. boreale, revealed the interspecific hybrids have naturally and repeatedly generated among the species and it makes to difficult and complicate to understand the evolutionary history in the genus.
ACKNOWLEDGMENTS
This study was supported by Basic Science Research Program through National Research Foundation of Korea (NRF) the funded by the Ministry of Education (No. 2017R1D1A1A02018573).
Fig. 1.
Map showing the localities of the populations which were described in the Table 1. ●, The diploid population (2 n = 18); ○, the tetraploid population (2 n = 36); ★ & ☆, The previously reported diploid populations. 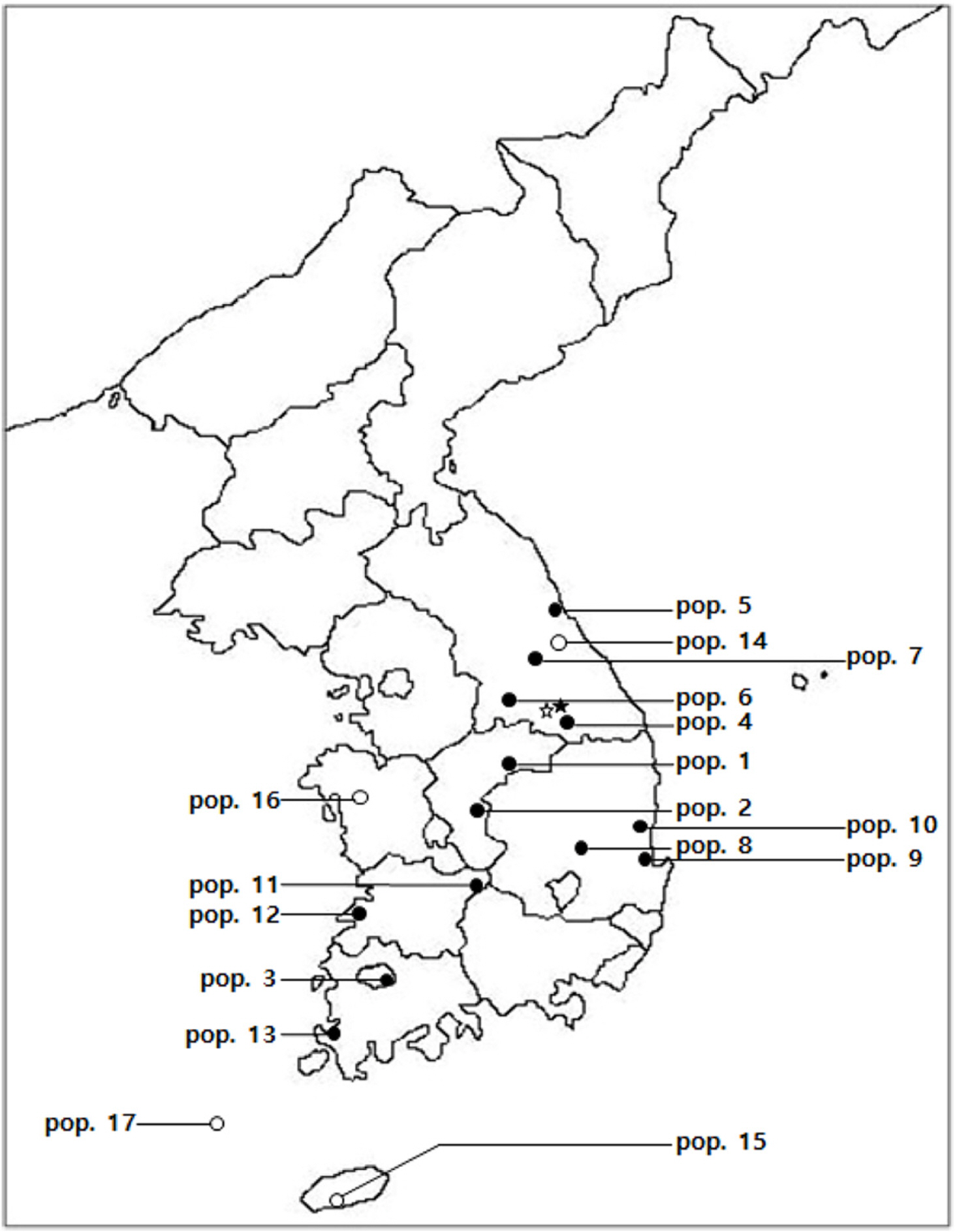
Fig. 2.
Representative leaf shapes of five populations of Chrysanthemum indicum. A. Pop. 1. B. Pop. 2. C. Pop. 3. D. Pop. 14. E. Pop. 15. Scale bars = 1 cm.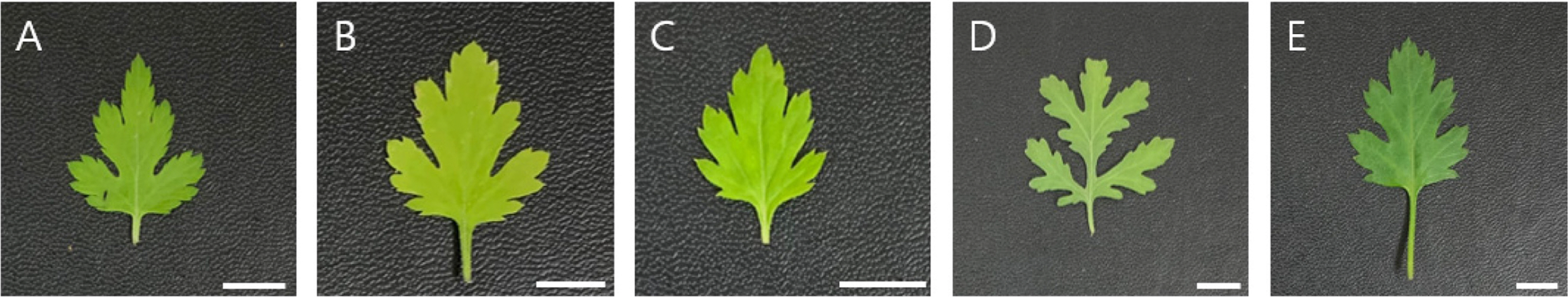 
Fig. 3.
Metaphase chromosomes during the mitosis of Chrysanthemum indicum. A. Pop. 1 (2n = 18). B. Pop. 2 (2n = 18). C. Pop. 3 (2n = 18). D. Pop. 14 (2n = 36). E. Pop. 15 (2n = 36). Scale bar = 10 μm. 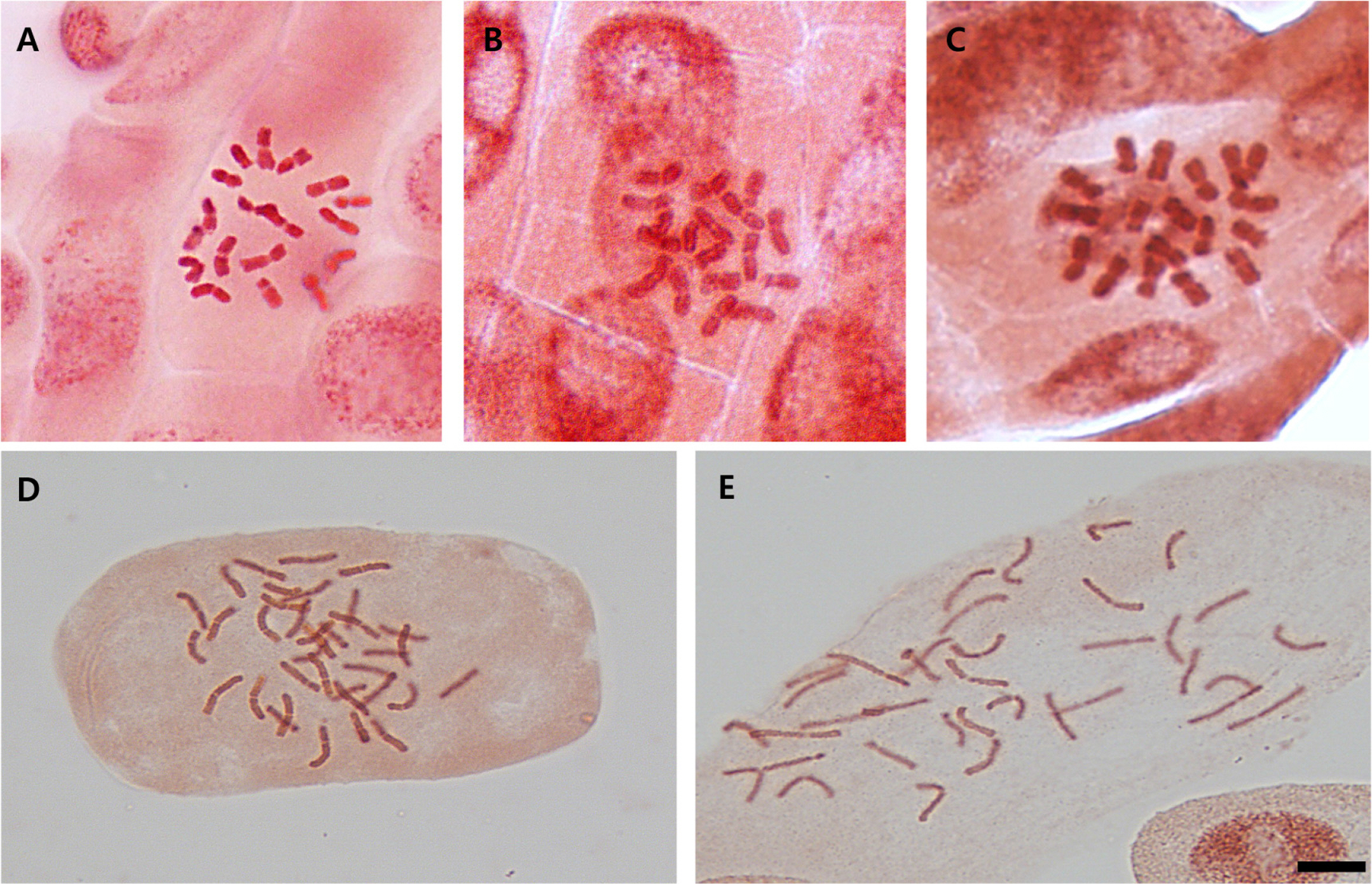
Table 1.
Collection sites information and chromosome number of Chrysanthemum indicum populations investigated in the study.
|
Pop. No. |
Locality |
Coordinate |
Elevation (m) |
Voucher specimen |
Chromosome number |
|
1 |
Songgye-ri, Hansu-myeon, Jecheon-si, Chungcheongbuk-do |
36.86°N, 128.08°E |
360 |
CBNU2018-0267 |
2x, 2n = 18 |
|
2 |
Songnisan-myeon, Boeun-gun, Chungcheongbuk-do |
36.53°N, 127.82°E |
338 |
CBNU2019-0225 |
2x, 2n = 18 |
|
3 |
Mudengsan, Geumgok-dong, Gwangju, Jeollanam-do |
35.13°N, 126.98°E |
520 |
CBNU2019-0226 |
2x, 2n = 18 |
|
4 |
Beagisan Mt., Nam-myeon, Jeongseon-gun, Gangwon-do |
37.17°N, 128.42°E |
380 |
CBNU2019-0227 |
2x, 2n = 18 |
|
5 |
Gwangjin-ri, Hyeonnam-myeon, Yangyang-gun, Gangwon-do |
37.96°N, 128.76°E |
65 |
CBNU2019-0209 |
2x, 2n = 18 |
|
6 |
Baekdal-ri, Ucheon-myeon, Hoengseong-gun, Gangwon-do |
37.44°N, 128.06°E |
220 |
CBNU2018-0206 |
2x, 2n = 18 |
|
7 |
Cheokcheon-ri, Jinbu-myeon, Pyeongchang-gun, Gangwon-do |
37.69°N, 128.50°E |
970 |
CBNU2018-0220 |
2x, 2n = 18 |
|
8 |
Binggyegyegok, Chunsan-myeon, Uiseong-gun, Gyeongsangbuk-do |
36.22°N, 128.75°E |
120 |
CBNU2019-0216 |
2x, 2n = 18 |
|
9 |
Josa-ri, Songna-myeon, Buk-gu, Pohang-si, Gyeongsangbuk-do |
36.22°N, 129.38°E |
28 |
CBNU2019-0214 |
2x, 2n = 18 |
|
10 |
Docheon-ri, Namjeong-myeon, Yeongdeok-gun, Gyeongsangbuk-do |
36.31°N, 129.35°E |
34 |
CBNU2019-0213 |
2x, 2n = 18 |
|
11 |
Deogyusan Mt., Seolcheon-myeon, Muju-gun, Jeollabuk-do |
35.88°N, 127.78°E |
717 |
CBNU2019-0221 |
2x, 2n = 18 |
|
12 |
Mapo-ri, Byeonsan-myeon, Buan-gun, Jeollabuk-do |
35.64°N, 126.47°E |
20 |
CBNU2019-0220 |
2x, 2n = 18 |
|
13 |
Yudalsan Mt., Mokwon-dong, Mokpo-si, Jeollanam-do |
34.78°N, 126.36°E |
32 |
CBNU2018-0396 |
2x, 2n = 18 |
|
14 |
Sangjinbu-ri, Jinbu-myeon, Pyeongchang-gun, Gangwon-do |
37.65°N, 128.57°E |
540 |
CBNU2018-0221 |
4x, 2n = 36 |
|
15 |
Cheonjeyeon, Jungmun-dong, Seogwipo-si, Jeju-do |
33.24°N, 126.42°E |
17 |
CBNU2018-0398 |
4x, 2n = 36 |
|
16 |
Yongbongsan Mt., Sangha-ri, Hongbuk-myeon, Hongseong-gun, Chungcheongnam-do |
36.64°N, 126.64°E |
139 |
CBNU2019-0197 |
4x, 2n = 36 |
|
17 |
Gageodo-gil, Heuksan-myeon, Sinan-gun, Jeollanam-do |
34.03°N, 125.07°E |
215 |
CBNU2019-0228 |
4x, 2n = 36 |
|
★ |
Kwangha-ri, Jeongseon-eup, Jeongseon-gun, Gangwon-do |
Previously reported diploid (2x, 2n = 18) population (Kim et al., 2003) |
|
☆ |
Baekun-ri, Mitan-myeon, Pyungchang-gun, Gangwon-do |
Literature Cited
Bailey, LH and Bailey, EZ. 1976. A concise dictionary of plants cultivated in the United States and Canada. Macmillan Publishing Company, New York, USA. 266-269.
Bremer, K and Humphries, CJ. 1993. Genetic monograph of the Asteraceae-Anthemideae. Bulletin of the Natural History Museum, Botany Series 23: 71-177.
Dowrick, GJ. 1952. The chromosome of Chrysanthemum, I: the species. Heredity 6: 365-375.   Du, B. Lin, Q. Zhu, C and Ke, S. 1989. Karyotype studies of two species of Dendranthema. Journal of Wuhan Botanical Research 7: 293-296.
Fu, L. Chen, T. Lang, K. Tao, H and Lin, Q. 2005. Higher Plants of China. 11: Qingdao Publishing House, Beijing. 341-348.
Grant, V. 1981. Plant Speciation. 2nd ed. Columbia University Press, New York. 432 pp.
Hotta, M. Yamagawa, N. Hirai, Y and Shiuchi, T. 1996. Taxonomical notes on plants of southern Japan III. Distribution and taxonomy of the Dendranthema ornatum group around southern Kyushu. Acta Phytotaxonomica et Geobotanica 47: 91-104.
Jeong, EH. 2011. Morphological characteristics and relationships of Korean native Chrysanthemum indicum collections. PhD dissertation,. Gyeongsang National University; Jinju, Korea. 113.
Kim, JS. Oginuma, K and Tobe, H. 2008. Analysis of meiotic chromosome behaviour in diploid individuals of Chrysanthemum zawadskii and related species (Asteraceae): evidence for chromosome rearrangements. Cytologia 73: 425-435.  Kim, JS. Pak, J-H. Seo, B-B and Tobe, H. 2003. Karyotypes of metaphase chromosomes in diploid populations of Dendranthema zawadskii and related species (Asteraceae) from Korea: diversity and evolutionary implications. Journal of Plant Research 116: 47-55.   Kim, JS. Pak, J-H and Tobe, H. 2004. Chromosome number of Dendranthema coreana (Asteraceae). Acta Phytotaxonomica et Geobotanica 55: 63-64.
Kim, JS and Tobe, H. 2009. Variations of leaf thickness in the Chrysanthemum zawadskii complex and in two related Korean species: C. boreale and C. indicum (Asteraceae). Korean Journal of Plant Taxonomy 39: 29-34.   Kitamura, S. 1940. Compositae Japonicae. Memoirs of the College of Science, Kyoto Imperial University, Series B Biology 15: 316-375.
Lee, YN. 1975. Taxonomic study on white flowered wild Chrysanthemum in Asia. Korean Journal of Botany 14: 63-73.
Lee, YN and Oh, YG. 1976. Taxonomic study on yellow-flowered wild Chrysanthemum in Korea. Journal of Korean Research Institute for Better Living 17: 143-154 (in Korean).
Lee, TB. 2003. Coloured Flora of Korea. Hyangmunsa, Seoul. 2096 (in Korean).
Lee, YN. 2006. New Flora of Korea II. Kyohaksa, Seoul. 318-325.
Nakata, M and Kumagai, A. 1999. Octoploid cytotype of Dendranthema zawadskii (Asteraceae) found in Iwate prefecture and its implications in evolutional history. Bulletin of the Botanical Garden of Toyama 4: 1-25.
Nakata, M and Tanaka, R. 1987. Species of Chrysanthemum in Japan in the study of chromosomes. . In Proceedings of the Sino-Japan Symposium on plant chromosomes research. Tanaka, R. Chen, R (eds.), Hiroshima. Nichiki, 17-22.
Nakata, M. Tanaka, R. Taniguchi, K and Shimotomai, N. 1987. Species of wild Chrysanthemum in Japan: cytological and cytogenetical views on its entity. Acta Phytotaxonomica et Geobotanica 38: 241-259.
Nicolson, DH. 1999. Report of the general committee: 8. Taxon 48: 373-378.  Oberprieler, CH. Vogt, R and Watson, LE. 2007. Tribe Anthemideae. In The Families and Genera of Vascular Plants. 8: Flowering Plants. Eudicots: Asterales. Kadereit, JW. Jeffrey, C (eds.), Springer, Berlin. 342-374.
Ohwi, J. 1984. Flora of Japan. Smithsonian Institution, Washington, DC. 889-893.
Shimotomai, N. Tanaka, R. Masumori, S and Ishiguro, M. 1956. Über die Polyploidie und geographische Verbreitung bei Chrysanthemum japonense Nakai. Botanical Magazine 69: 514-518.  Song, HS. Kim, SM and Park, YJ. 2012. Comparison of vegetation and habitat condition of Dendranthema boreale and Dendranthema indicum in Korea. Korean Journal of Medicinal Crop Science 20: 20-26.   Tanaka, R. 1959. On the speciation and karyotypes in diploid and tetraploid species of Chrysanthemum boreale×Ch. makinoi. Journal of Science of the Hiroshima University, Series B, Division 2, Botany 9: 31-40.
Tanaka, R. 1960. On the speciation and karyotypes in diploid and tetraploid species of Chrysanthemum. V. Chrysanthemum× yoshinaganthum (2n=36). Cytologia 25: 43-58.  Taniguchi, K. 1987. Cytogenetical studies on the speciation of tetraploid Chrysanthemum indicum L. with special reference to C-bands. Journal of Science of the Hiroshima University, Series B, Division 2, Botany 21: 105-157.
Trehane, P. 1995. Proposal to conserve Chrysanthemum L., with a conserved type (Compositae). Taxon 44: 439-441.  Tsukaya, H. 2002. Leaf anatomy of a reophyte, Chrysanthemum yoshinaganthum (Asteraceae), and of hybrids between D. yoshinaganthum and a closely related nonreophyte, D. indicum. Journal of Plant Research 115: 329-333.   Zhao, HE. Liu, ZH. Hu, X. Yin, JL. Li, W. Rao, GY. Zhang, XH. Huang, CL. Anderson, N. Zhang, QX and Chen, JY. 2009. Chrysanthemum genetic resources and related genera of Chrysanthemum collected in China. Genetic Resources and Crop Evolution 56: 937-946.  
|
|














